Onichtchouk Lab
PD rer. nat. Daria Onichtchouk
Research project
I am interested in the regulatory mechanisms which underlie early embryonic development, and using zebrafish model system to investigate this question. The key point at the beginning of development of any multicellular organism is so called called maternal-to-zygotic transition, or Midblasula Transition (MZT, or MBT), the short time period when subset of maternal RNAs rapidly degrades, and transcription of zygotic genes commences. After MBT in vertebrates, embryonic cells are pluripotent for certain time period, and later become progressively restricted to specific developmental fates, according to position-specific signals they recieve from embryo environment. Start of transcription of the first zygotic gene cohort is precisely synchronized by MBT. Very little is known about mechanisms which specifically determine the set of genes to be transcribed at MBT, as well as about genetic control of subsequent transition from pluripotency to lineage restriction in the embryonic cells.
Previously, we demonstrated that the deficiency for transcription factor Pou5f1 results in global temporal shifts in gene expression during early development (Onichtchouk etal., 2010). Recently, we have shown that Pou5f1 together with SoxB1 transcription factors (Sox2, Sox3, Sox19a and Sox19b) specifically activate the first zygotic gene cohort in zebrafish, expressed at MBT (Leichsenring et al., 2013, submitted). Pou5f1 and SoxB1 act by binding the composite SOX-POU binding sites, abundant in the promoters and enhancers of the first zygotic genes, encoding predominantly developmental regulators (Fig1).
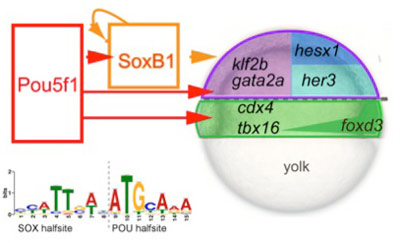
Mammalian orthologues of Pou5f1 (Oct4) and Sox2 are the key transcription factors, which define indefinite pluripotency in mammalian ES cells. Identity of the binding sites and orthology of the target gene sets of Pou5f1 and SoxB1 between zebrafish and mammals suggests that early gene regulatory network in vertebrates has common evolutionary origin.
Based on the previous work, we aim to further explore the link between pluripotency and zygotic gene activation in zebrafish embryo. Characterization of Pou5f1 and SoxB1 regulatory network will involve genetics, genomics and molecular biological approaches and focus on several projects:
- Epigenetics of zebrafish Midblastula Transition: connection with Pou5f1 and SoxB1 transcriptional regulators BIOSS
- Functional roles of miRNA targets of Pou5f1 and SoxB1
- Characterisation of FoxD3 and FoxD5 transcription factors
Interested in Master and Bachelor thesis are wellcome!
Please, contact directly :
Daria Onichtchouk
Bio 1, Raum 2042
Daria.Onichtchouk@biologie.uni-freiburg.de
Tel.: ++49 / 761 / 203 - 2595


